First Detection of Xylella fastidiosa Infecting Cherry (Prunus avium) and Polygala myrtifolia Plants, in Mallorca Island, Spain
Authors
D. Olmo (1), A. Nieto (1), F. Adrover (1), A. Urbano (2), O. Beidas (3), A. Juan (3), E. Marco-Noales (4), M. M. López (4), I. Navarro (4), A. Monterde (4), M. Montes-Borrego (5), J. A. Navas-Cortés (5), B. B. Landa (5).
Affiliations
(1) Serveis de Millora Agrària, Govern Balear, 07009, Palma de Mallorca, Spain.
(2) TRAGSA, Empresa de Transformación Agraria, Delegación de Baleares, 07009, Palma, Spain.
(3) Servicio de Agricultura, Conselleria de Medi Ambient, Agricultura i Pesca, 07006 Palma, Spain.
(4) Instituto Valenciano de Investigaciones Agrarias (IVIA), 46113 Moncada, Valencia, Spain.
(5) Instituto de Agricultura Sostenible (IAS-CSIC), 14004, Córdoba, Spain.
Disease Note
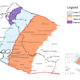
Xylella fastidiosa, a quarantine organism in the European Union (EU), causes diseases in a wide variety of plants such as almond, cherry, grape, citrus, elm, olive, and coffee trees and many ornamental plants. Since the detection of the bacterium in Italy (2013), where it is associated to a severe epidemic on olive trees, the pathogen has also been detected in France (2015) and Germany (2016) (EPPO 2016). Due to the recent outbreaks and to different interceptions, the EU has implemented annual surveys in its member states to prevent new introductions or the spread of this harmful organism. During official surveys in late autumn 2016 in Mallorca Island, Spain, some cherry and Polygala myrtifolia plants located in a garden center near the locality of Manacor showed symptoms of marginal leaf scorch, leaf chlorosis, defoliation, and general decay. DNA was extracted from leaf veins and petioles of symptomatic leaves using the CTAB extraction method (EPPO 2016). DNA extracts were tested for the presence of X. fastidiosa by real-time PCR using two species-specific protocols (EPPO 2016): primers XF-F/XF-R and a dual-labeled probe XF-P (Harper et al. 2010, erratum 2013) and primers HL5/HL6 and Taqman probe (Francis et al. 2008). DNA from X. fastidiosa subsp. pauca strain CoDiRo or subsp. fastidiosa strain Temecula were used as positive controls. Three out of four cherry trees and four out of seven P. myrtifolia plants showing symptoms gave positive real-time amplification. These preliminary results were further confirmed by conventional PCR assays, using species specific primers RST31/RST33 (Minsavage et al. 1994), which yielded the expected amplicon of ∼700 bp of the RNA polymerase sigma 70 factor. Attempts to culture isolates from petioles tissues on PD2 and BCYE media failed for the P. myrtifolia plants. On the contrary, bacterial colonies typical of X. fastidiosa were recovered from one cherry plant after plate incubation for 14 days at 28°C. Colonies were suspended in water and boiled 10 min to obtain DNA before PCR analysis. Sequence analysis of the amplicon of the sigma 70 factor from all positive samples showed 95 to 100% identity with other X. fastidiosa sequences from GenBank, which confirmed the presence of X. fastidiosa DNA in the plants. Additionally, seven housekeeping genes (Yuan et al. 2010) were amplified using an annealing temperature of 58°C from both extracted plant or bacterial DNAs to identify the subspecies and sequence type (ST) of X. fastidiosa. Based on this MLST analysis, the ST1, related to X. fastidiosa subsp. fastidiosa, was found on one cherry tree and three P. myrtifolia plants. These four plants shared 100% nucleotide sequence identity for the sigma 70 factor (MF401540). Surprisingly, the fourth P. myrtifolia sample presented sequences that simultaneously have been only described in X. fastidiosa subsp. multiplex isolates (i.e., leuA_3, holC_3, nuoL_3, and gltT_3), although only four alleles could be amplified. This plant sample showed a sigma 70 factor sequence (MF401541) identical to that of X. fastidiosa subsp. multiplex isolates present in GenBank and different to those of samples infected by X. fastidiosa subsp. fastidiosa ST1. To our knowledge, this is the first detection of X. fastidiosa in Spain and of the subsp. fastidiosa on cherry and P. myrtifolia in Europe, underlining the presence of different genotypes in this island and the risk of introducing additional genetic diversity for X. fastidiosa to Europe. Eradication measures have been taken in the garden center according to the Spanish contingency plan and EU legislation.
The work was supported by funding from the European Union’s Horizon 2020 research and innovation programme under grant agreement No 635646 POnTE (Pest Organisms Threatening Europe) and No 727987 XF-ACTORS (Xylella Fastidiosa Active Containment Through a multidisciplinary-Oriented Research Strategy), and the Spanish Ministry of Agriculture and Fisheries, Food and Enviroment (MAPAMA).
Published on October, 2017 by PLANT DISEASE.